Graphical Representation of Biomolecules
Protein structure
There are various ways to depict the tertiary structure of a protein, and a choice can be made to highlight whichever of its molecular features is the most relevant. The molecular models in these pages are generated using JSmol, which follows the standard practice of depicting sections of alpha-helix as ribbons, and sections of beta-sheet as flattened arrows. The trace view is the simplest option for showing secondary structure, limiting the representation to a smooth line that follows the path of the polypeptide N–C–C backbone.
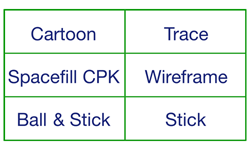
MOUSEOVER the frame below for various views of the protein crambin
All views show the same molecule in the same orientation
See below for an interactive model of crambin, with the molecule in the same orientation as depicted in these graphics.
Interactive Examples
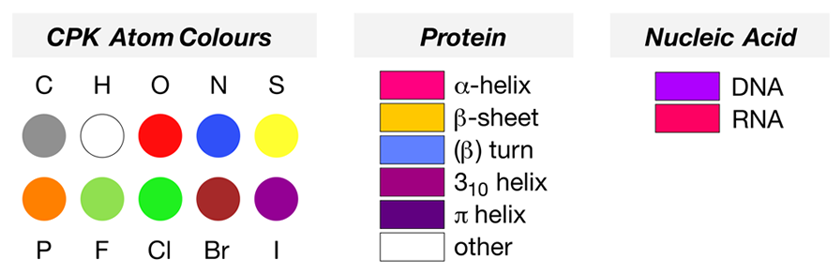
The simple protein crambin (1CRN) contains sections of α-helix, represented in the cartoon mode as magenta ribbons, and β-sheet, represented as flat yellow arrows indicating the N-to-C-to-C direction of the residues involved. The structure 1D66 is a protein-DNA complex, which has sections of protein configured as α-helix (magenta ribbons) as well as a DNA base pair sequence (the double helix coloured in electric purple).
Water structure of a hydrophobic protein at atomic resolution: Pentagon rings of water molecules in crystals of crambin.
[PDB structure code 1CRN]; M. M. Teeter, Proc. Natl. Acad. Sci. USA, 1984, 81, 6014–6018
DNA recognition by GAL4: structure of a protein-DNA complex. [PDB structure code 1D66]
R. Marmorstein, M. Carey, M. Ptashne and S. C. Harrison, Nature, 1992, 356, 408–414
